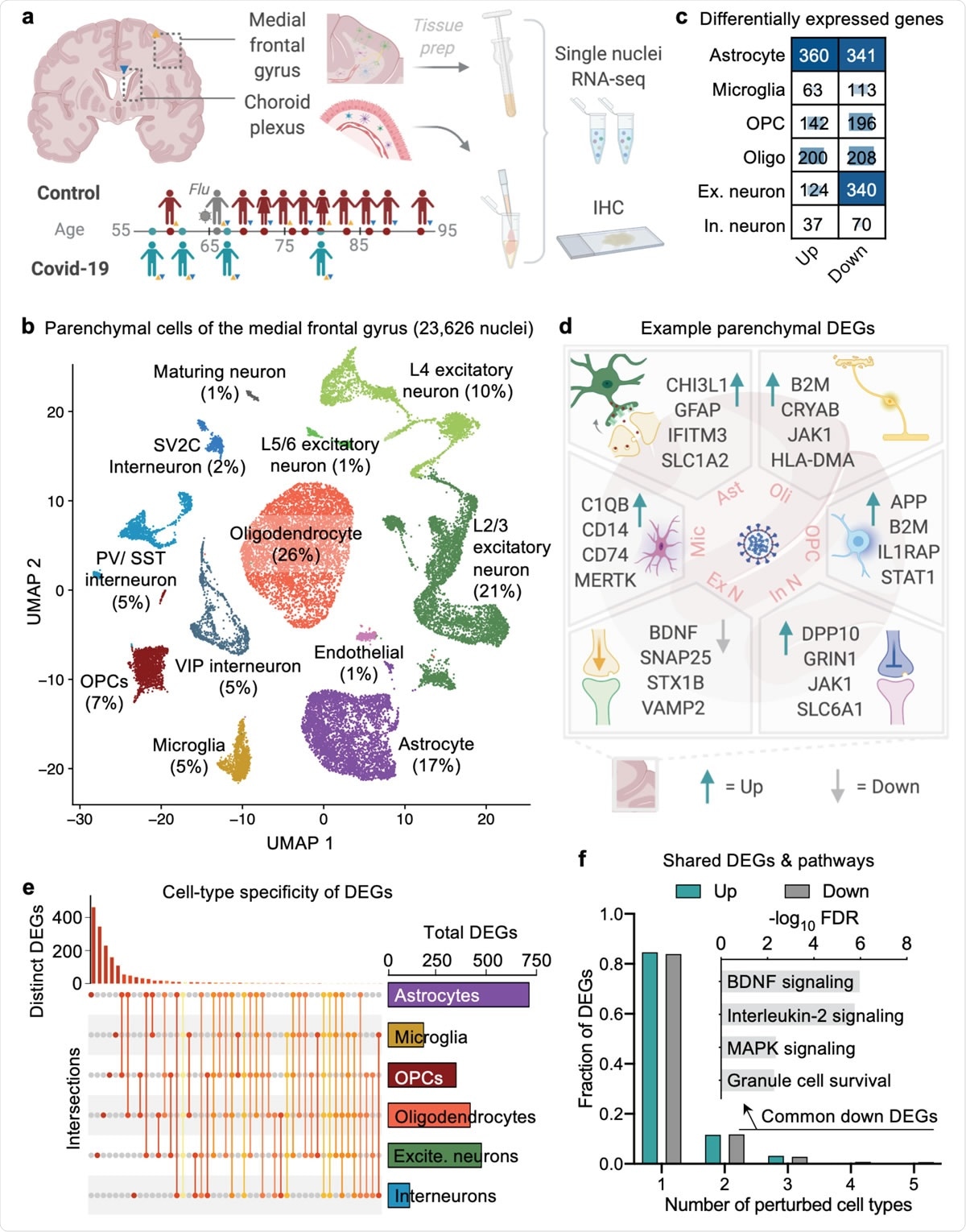
Systems-Level Properties of EGFR-RAS-ERK Signaling Amplify Local Signals to Generate Dynamic Gene Expression Heterogeneity - ScienceDirect
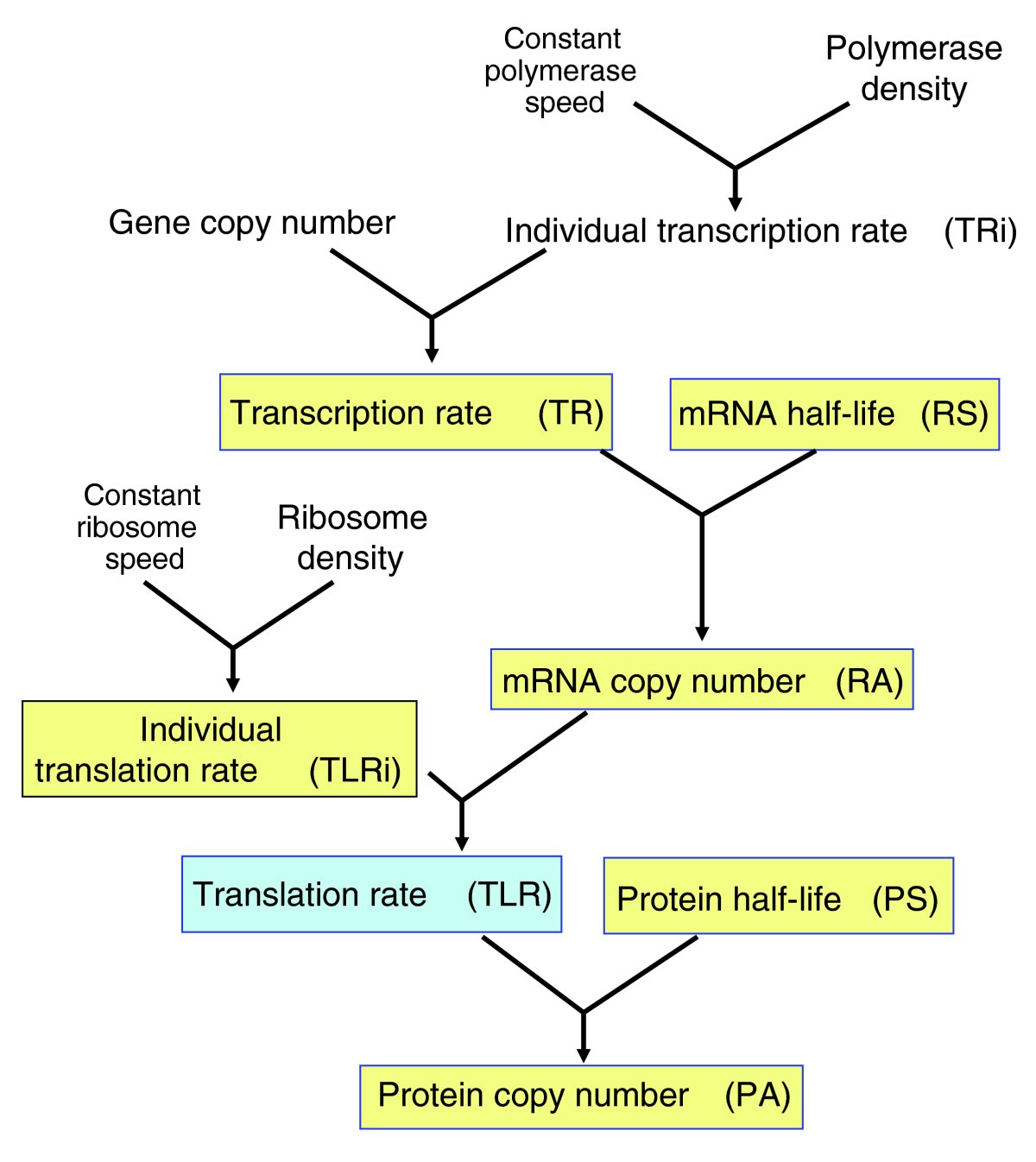
Common gene expression strategies revealed by genome-wide analysis in yeast | Genome Biology | Full Text
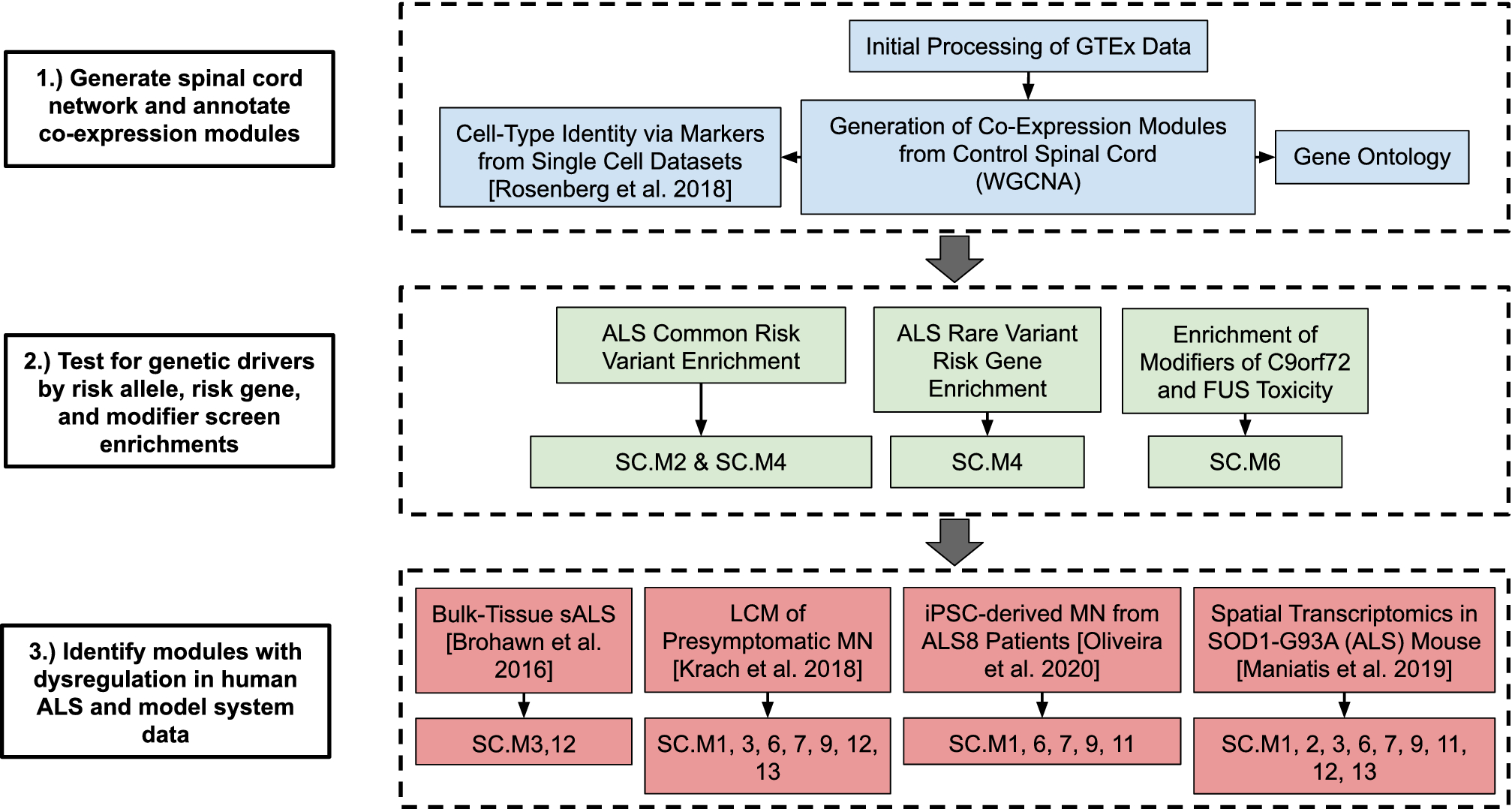
Gene co-expression network analysis in human spinal cord highlights mechanisms underlying amyotrophic lateral sclerosis susceptibility | Scientific Reports
Predicting Spatial and Temporal Gene Expression Using an Integrative Model of Transcription Factor Occupancy and Chromatin State | PLOS Computational Biology
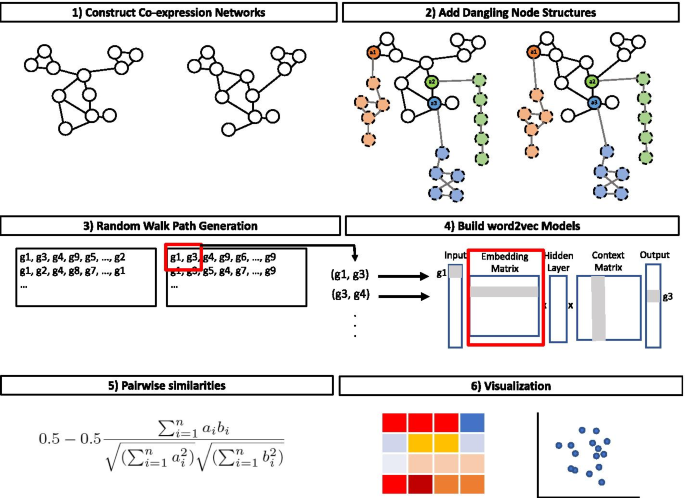
Juxtapose: a gene-embedding approach for comparing co-expression networks | BMC Bioinformatics | Full Text

Overview of methods and tools used to create and analyse co-expression networks constructed from gene expression data | RNA-Seq Blog

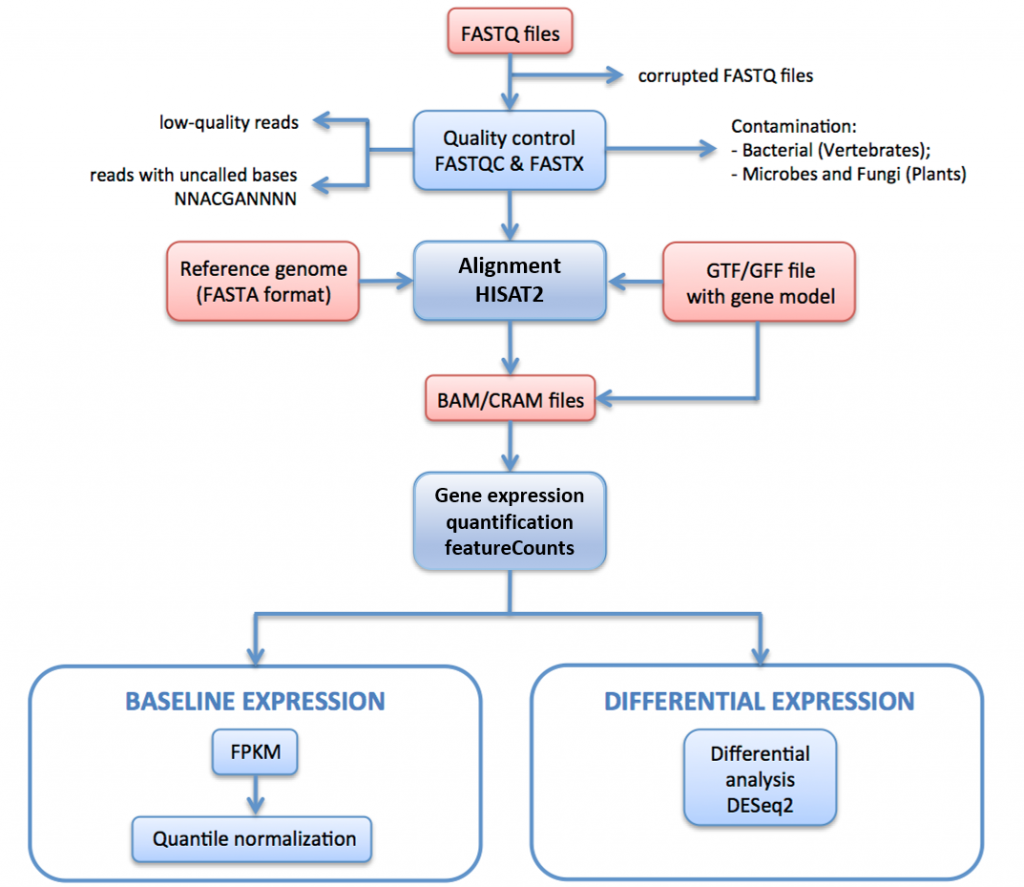
Flow-chart showing the procedure to generate gene expression profiles.... | Download Scientific Diagram
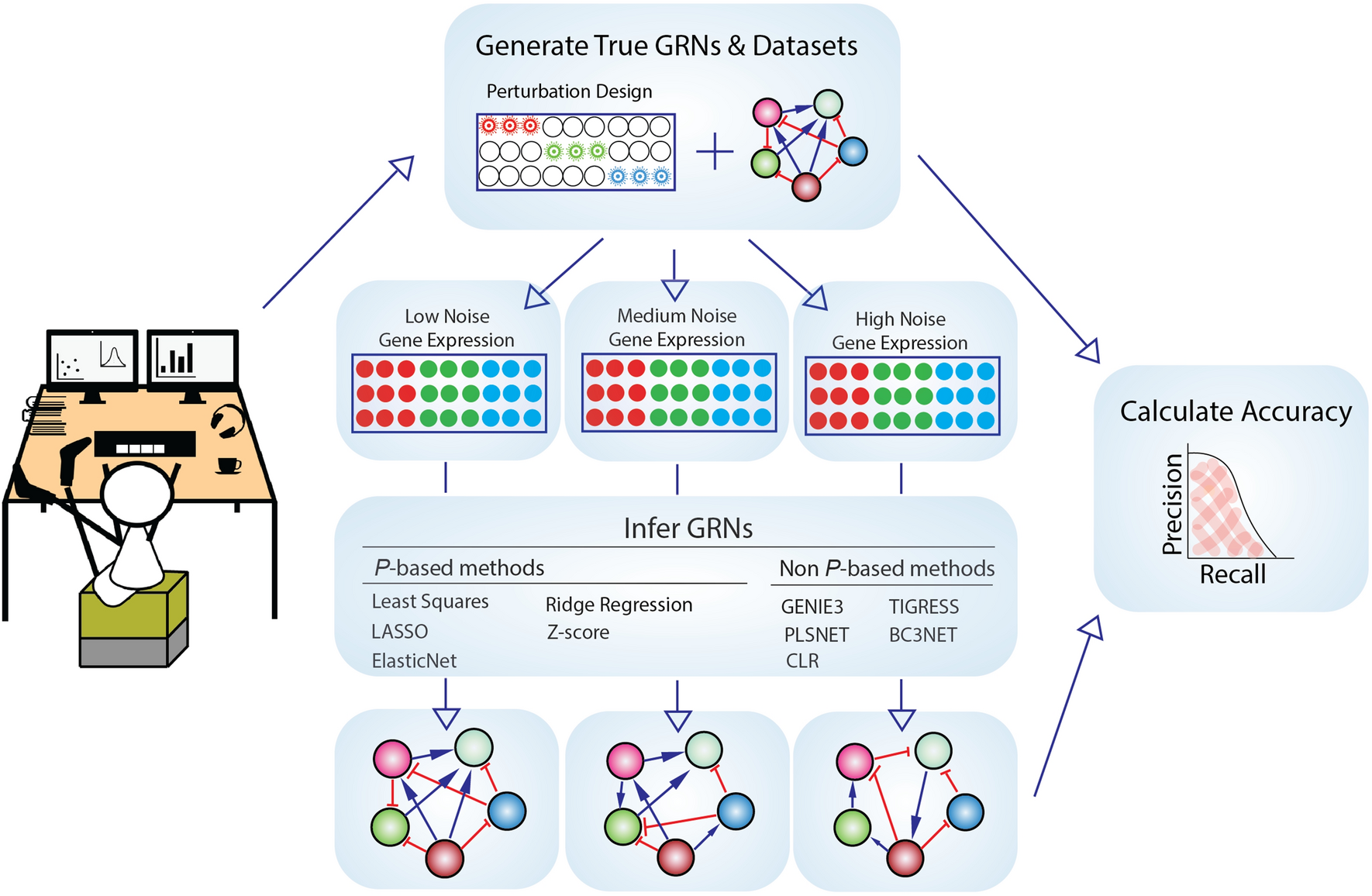
Knowledge of the perturbation design is essential for accurate gene regulatory network inference | Scientific Reports